In the last year, SQL has wound its way deeper and deeper into my R workflow. I switch between the two every day, but up to now, I’ve been slow diving into the SQL tools RStudio provides. This post documents the little tricks I’ve gathered scouring RStudio Community posts, blogposts, and other resources for help upgrading my R-SQL workflow. To see just the code, take a look at the associated GitHub gist.
My starting point
My typical workflow currently looks like this:
- Draft and refine SQL queries in the Rubymine IDE
- Bring the data into R/RStudio by calling
DBI::dbGetQuery()or a similar function - Do complex manipulations or visualizations of the data in R/RStudio
- Iterate on the steps above
This workflow is something I’ve largely absorbed from my coworkers. Some of the peculiarities are due to historical accidents – for example, many of the early developers at the company used Ruby, so RubyMine also became the IDE of choice for working with SQL. That said, this flow works pretty well for us.
Over time, I found my own reasons for using RubyMine (or really, any other IDE with advanced SQL support):
- easy to kill queries
- syntax highlighting (code coloring) for SQL
- auto-completion for SQL
- auto-formatting for SQL (cmd + option + l is my favorite shortcut ever in RubyMine)
- helpful hints/errors when something’s wrong with my query

I’m glad I was forced to venture out of my RStudio comfort zone, but I’ve begun to discover that some of the features I love about RubyMine actually exist in RStudio as well.
Previewing SQL in RStudio
Up to now, I’d only used dataframe-style previews of query results, such as in the example below:
library(dplyr)
library(dbplyr)
conn <- src_memdb() # create a SQLite database in memory
copy_to(conn,
storms, # this is a dataset built into dplyr
overwrite = TRUE)
tbl(conn, sql("SELECT * FROM storms LIMIT 5"))# Source: SQL [?? x 13]
# Database: sqlite 3.30.1 [:memory:]
name year month day hour lat long status category wind pressure
<chr> <dbl> <dbl> <int> <dbl> <dbl> <dbl> <chr> <chr> <int> <int>
1 Amy 1975 6 27 0 27.5 -79 tropi… -1 25 1013
2 Amy 1975 6 27 6 28.5 -79 tropi… -1 25 1013
3 Amy 1975 6 27 12 29.5 -79 tropi… -1 25 1013
4 Amy 1975 6 27 18 30.5 -79 tropi… -1 25 1013
5 Amy 1975 6 28 0 31.5 -78.8 tropi… -1 25 1012
# … with 2 more variables: ts_diameter <dbl>, hu_diameter <dbl>It turns out that within RStudio, there are actually (at least) two other methods of previewing SQL queries. (Thank you to RStudio Community posts such as this one for enlightening me.) These two methods also have the added benefit of SQL code with syntax highlighting.
1. Preview a .sql file
When you open a new .sql file in RStudio, it automatically populates the file with the following code:
-- !preview conn=DBI::dbConnect(RSQLite::SQLite())
SELECT 1I’d always blindly deleted the header, but it turns out that all this time, RStudio had been trying to tell me that I can actually preview queries in RStudio! If you supply it with a working database connection (no spaces around the =!) and click the Preview button, a SQL Results tab pops up next to the Console with the query results.
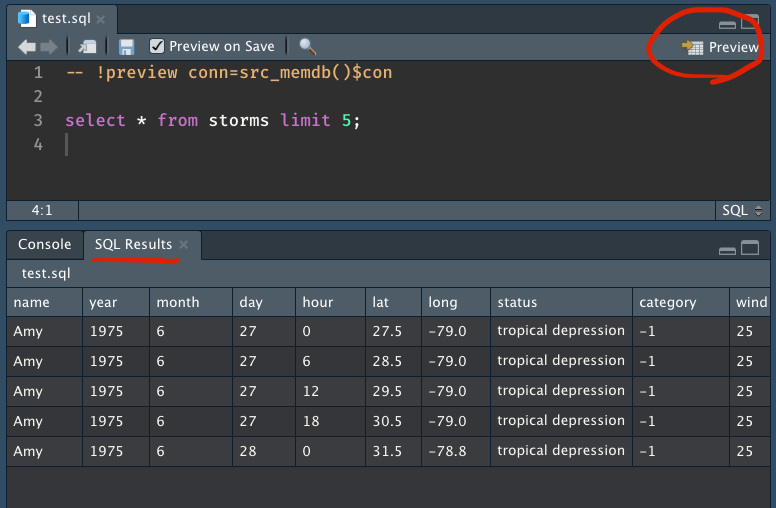
If you’ve run the previous dplyr/dbplyr code chunk in the same session, you’ll have both the connection and the data you need to test out this preview function yourself! Just create a .sql file with the following code:
-- !preview conn=src_memdb()$con
SELECT * FROM storms LIMIT 5
Note that unlike in the tbl() example above, you have to pull out the connection from the src_memdb() using $con. This seems to be a special case; other connections seem to work without extra fussing. For the rest of this post, I’ll refer to this connection as con with one “n” (with the exception of the gif below, where I gave it the lengthier name conn_isolated).
con <- src_memdb()$con2. SQL chunks in RMarkdown
I generally prefer to show RMarkdown output in the console 1 (and it looks like I’m not the only one). This means that when I run code in an .Rmd file, it feels more or less the same as when I run an .R file: the plots show up in the plots pane, code is run in the console, and so on.
While you can use SQL chunks with this setting, there is NO chunk preview option. You must trust your queries and knit the file to make sure everything runs. You get the syntax highlighting razzle-dazzle but alas– no preview.
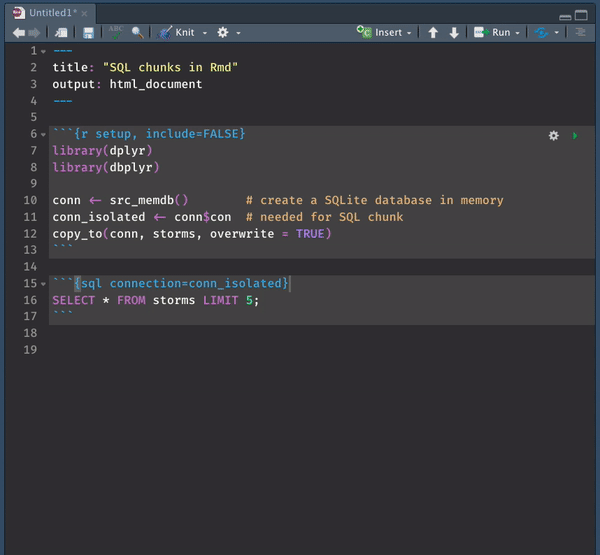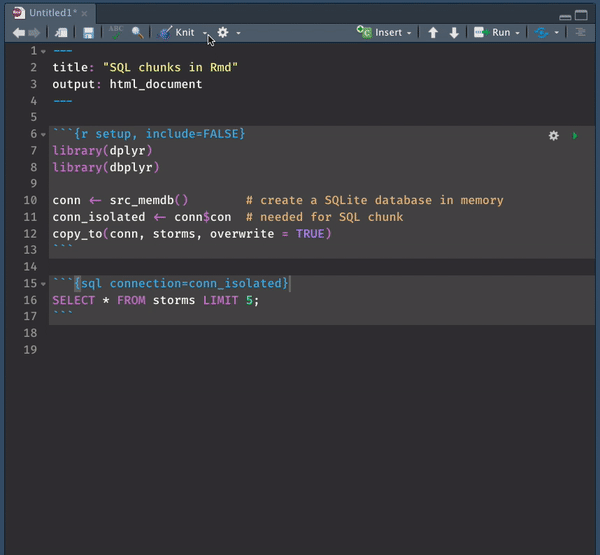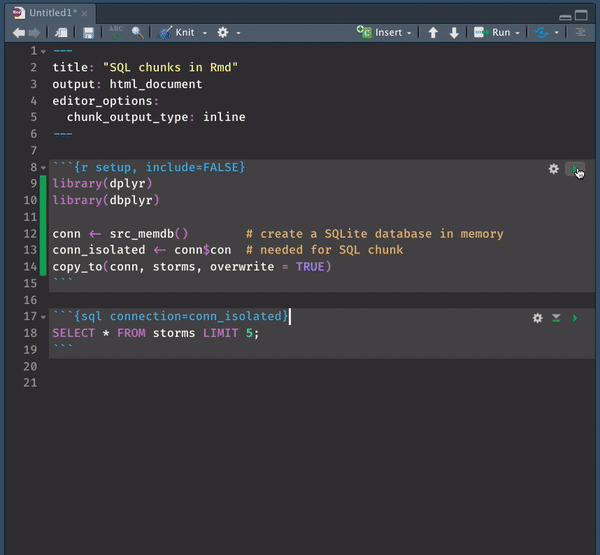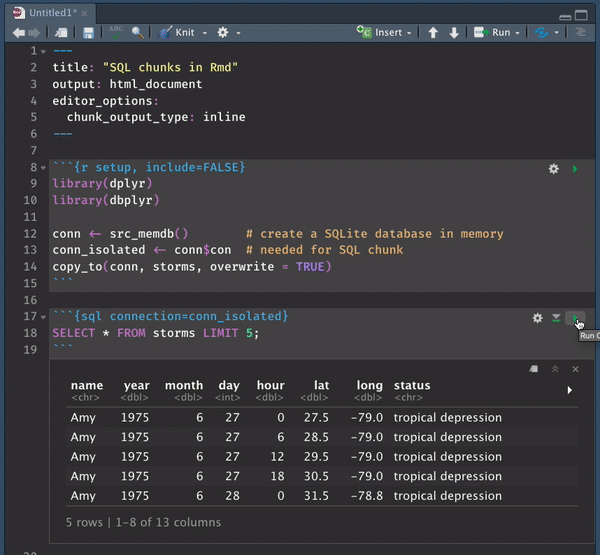
It is in this very specific case where inline mode wins big time. SQL previews magically become an option, allowing you to interact with your beautifully colored SQL code.
Passing variables to/from SQL chunks
When mixing R and SQL, you often want to (a) get data from a database and then continue manipulating it in R, or (b) modify the query by passing in variables from R.
SQL output as a variable
When using SQL chunks, you can specify an output variable using the output.var chunk option with the variable name as a string. 2
In inline mode, the preview will no longer appear when running the SQL chunk, but the variable will appear in your environment. Think of it like a regular variable assignment in R, where no output is printed when a variable is assigned (storm_preview <- "blah").3
```{sql connection=con, output.var="storm_preview"}
SELECT * FROM storms LIMIT 5;
``````{r}
storm_preview
```Providing query parameters
Adding a single variable is quite simple. Add ? before the variable in the SQL chunk and just make sure that the R variable has the same name.
```{r}
storm_status <- "hurricane"
``````{sql connection=con}
SELECT * FROM storms WHERE status = ?storm_status LIMIT 5;
```Sometimes, you cannot simply pass in a parameter. The glue_sql() function is great at managing some of the magic for you. To pass in a column name as a parameter, for example, you can just wrap the string in glue_sql() and it just works.
```{r}
col_name <- glue::glue_sql("status")
``````{sql connection=con}
SELECT * FROM storms WHERE ?col_name = 'hurricane' LIMIT 5;
```Note: Generally, you will also need to specify the .con (connection) so that glue will know what type of SQL syntax you need.
Now, what if you want to provide multiple parameters? Adding an * to the end of the variable collapses the vector into a single comma-separated expression, rather than outputting a vector of SQL expressions. This behavior is somewhat reminiscent of args and kwargs in Python, which uses * as an unpacking operator.
types <- c("hurricane", "tropical depression")
# without an asterisk
glue::glue_sql("{types}", .con = con)<SQL> 'hurricane'
<SQL> 'tropical depression'# with an asterisk
glue::glue_sql("{types*}", .con = con)<SQL> 'hurricane', 'tropical depression'Of course, to pass these parameters to our SQL chunk, we need to make sure we have the appropriate parentheses around ?storm_status for the IN to run.
```{r}
types <- c("hurricane", "tropical depression")
storm_status <- glue::glue_sql("{types*}", .con = con)
``````{sql connection=con}
SELECT * FROM storms WHERE status IN (?storm_status) LIMIT 5;
```SQL files meet chunks
What if you have a gigantic SQL query that you want to store in a separate file but you also want to use chunks?
I had no idea that chunks had a code parameter until I came across Christophe Dervieux’s comments in this RStudio Community discussion. You can use it read in a SQL file with your query, parameters and all.
```{sql connection=con, code=readLines("storm.sql"), output.var="storm_preview"}
``````{r}
storm_preview
```When you knit the file, the imported SQL (and its comments) is included in the output by default.
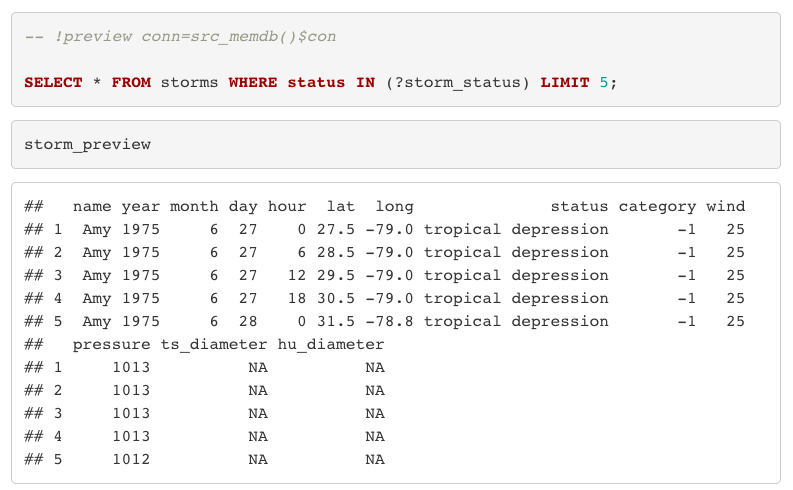
R & SQL – working hand-in-hand
At the end of the day, both R and SQL are–and will remain–big parts of my analysis flow, so it’s these little tricks that help to smooth out the little points of friction on the way. If you have more tricks to share with me, leave a comment or ping me on Twitter!

Creds to Allison Horst for the fun arms & faces!
For “console mode” Rmd’s:
Global Options–>R Markdown–> uncheckShow output inline for all R Markdown documents↩There are a few additional chunk option examples in the SQL language engine section of R Markdown: The Definitive Guide↩
For all my enthusiasm for syntax highlighting, I admit I do not understand how to do it properly when quoting RMarkdown chunks, though this RStudio blogpost gave me some clues. For more details on how to use syntax highlighting with blogdown, I recommend posts by Amber Thomas and Maëlle Salmon↩